In Post 1 I walked through the seven preprocessing stages from raw EEG to the 40x40 input the model sees. Stage 5 mentioned a corner-mask in passing. This post is what’s underneath that mention.
If you took the unmasked 40x40 topomap straight into a recon loss or a clustering metric, you’d be measuring agreement on a ring of pixels that contain no brain activity. The post argues that’s the wrong thing to do, shows the geometry, and gives the helper anyone touching this pipeline should use.
Where the corners come from
The 40x40 topomap is produced by scipy.griddata interpolating channel values onto a regular grid using the 2D-projected electrode positions. Electrodes only cover the inscribed head disc; the four corners of the square grid are outside that disc. griddata fills those pixels with whatever fill_value you pass it. The pipeline passes fill_value=0.
# process_eeg_signals.py: create_topographic_map
gy, gx = np.mgrid[ymin:ymax:TOPO_GRID*1j, xmin:xmax:TOPO_GRID*1j]
topo = griddata(pos_2d, vals, (gx, gy), method="cubic", fill_value=0)So every 40x40 topomap shipped through the pipeline has the same constant-zero ring of corner pixels. By construction. Same shape across participants, sessions, and cluster centroids. They look like part of the image but they’re not measurements of anything.
The mask, in one picture
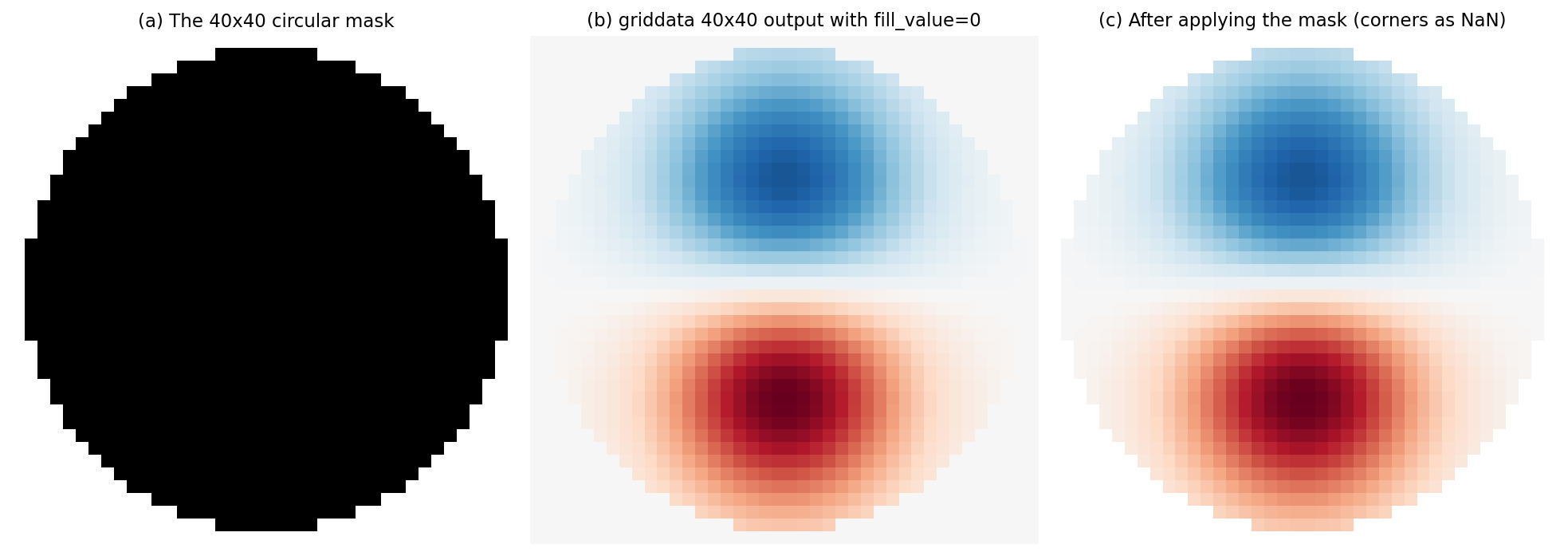
(a) The binary mask. White is inside the head disc, black is the padded ring. (b) A representative 40x40 topomap straight out of griddata. Notice the constant near-zero border outside the disc. (c) The same topomap after applying the mask. Corner pixels become NaN and are excluded from any per-pixel statistic.
Two numbers worth knowing:
- Pure geometry. A perfect inscribed circle in a 40x40 square covers 78.6% of the pixels. The corners are 21.4%.
- Our actual mask. We use
radius_factor=0.95to stay clear of edge artefacts from the cubic interpolation. With that margin, the masked ring is about 30% of the pixels.
The README rounds the geometric number to “~22%“. The mask in the code rounds the project number to ~30%. Both are correct, they’re answering different questions.
What breaks if you skip the mask
Every loss, per-pixel metric, and pairwise distance that touches the 40x40 image inherits the corner pixels. Each one fails in a slightly different way.
- Reconstruction loss wastes ~30% of its capacity. MSE on flat 1600-D tensors sums squared errors across all pixels and divides by 1600. The corner contributions are tiny (both ground-truth and reconstruction are near zero) but the denominator counts them, so the mean is artificially small. The model learns “agree with zero in the corners”, which is free.
- SSIM is inflated. SSIM slides a window across the image and measures local similarity. A window centred over a uniform-zero corner region matches perfectly between any two topomaps, because uniform vs uniform is the easy case for the metric. SSIM averages over windows, so the corner agreement props the score up.
- Pearson correlation between centroids is damped. Pearson centres each variable by subtracting its mean, then divides by the product of standard deviations. Constant pixels widen the variance estimate without contributing variance, deflating the denominator. Centroid correlations that should be near 1.0 come out lower than they really are.
- Silhouette, Davies-Bouldin, Calinski-Harabasz, Dunn all bias the same way. Each cluster-validity score uses centroid-to-centroid or point-to-point distances. Every pair of centroids agrees on the corner pixels. That’s a free chunk of “compactness” and “agreement” that shouldn’t be in the bookkeeping.
The fix in all four cases is the same: exclude the corners before the statistic.
The canonical helpers
mask_utils.py builds the mask once per process lifetime and returns whichever shape you need:
from mask_utils import (
get_topomap_mask_flat_torch_float, # (1600,) float, for masked MSE on (B, 1600) tensors
get_topomap_mask_flat_torch, # (1600,) bool, for column-indexing flat tensors
get_topomap_mask_2d_np, # (40, 40) bool, for SSIM masked-average + viz
get_topomap_mask_flat_np, # (1600,) bool, for sklearn / pycrostates pre-slicing
)The mask is a pure function of (H, W, radius_factor). No data dependency, no randomness, no per-batch state. It’s cached with lru_cache so any helper called a million times in training is reading from a tiny lookup, not rebuilding.
If you write a new metric or loss that consumes a 40x40 topomap, import the right helper and slice or multiply before reducing. One line.
What stays unmasked
Not every metric in the project touches the 40x40 image. The latent-space ones never do.
- Q1 (latent Euclidean), Q2 (latent |1/r|-1 ), KLD, polarity-invariance loss. All of these work on the encoder’s output vector in the latent space (size 32 by default). The corners of the input image don’t survive the encoder. No mask needed.
The convention is: mask only for things that look at the image. Anything that looks at the embedding is already corner-free by construction.
For people downloading the dataset
The masking convention applies to any consumer of the LEMON 40x40 topographic-map archive on Backblaze B2, not just code inside this project. There’s a runnable example at b2_docs/download_examples/apply_topomap_mask.py (in the public repo, same masking logic as mask_utils.py) with copy-pasteable NumPy and PyTorch snippets. If you’re training your own model on the cached .npy arrays, import either pair of helpers and stay consistent with radius_factor=0.95.
Why this is worth a post
Most pipelines that publish a “40x40 EEG topomap” archive document the format and stop there. The dead-corner question is one of those things you only discover when you compute a metric, get a suspicious number, dig in, and realise 30% of your image is uniform zeros. Posting it as a convention means anyone using the data starts on the right side of the wall.
Next post: 4-quadrant evaluation. Why one cluster score is never enough for EEG, and what the {latent, decoded} x {Euclidean, |1/r|-1} grid actually tells you.